What biologists call a species is becoming more than just a name
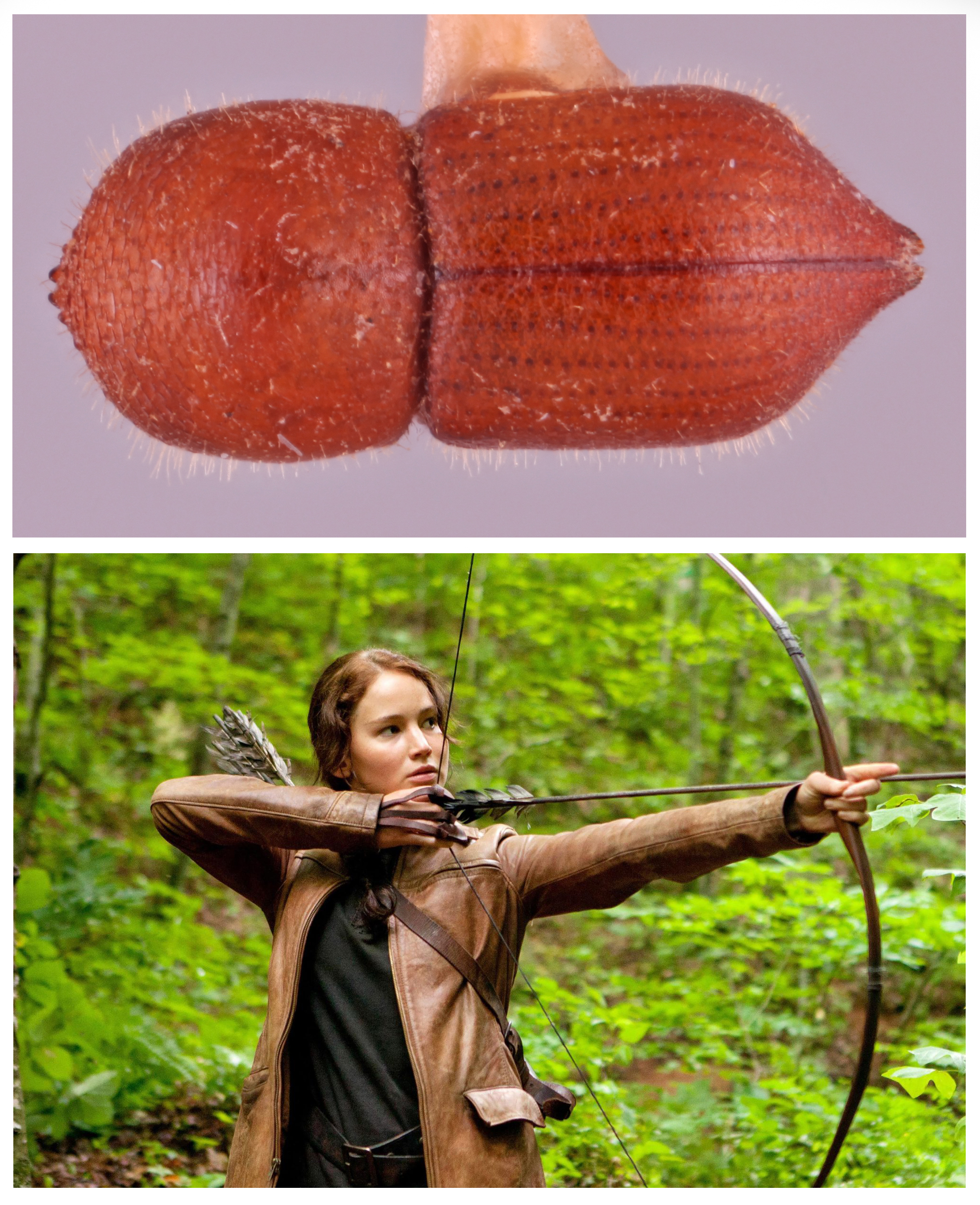

Table of Contents
You may not be familiar with the scientific names of ambrosia beetles. These insects belong to the genus Coptoborus. But you are likely to recognize some iconic female characters from science fiction among a few of their species names. Consider C. katniss, C. scully and C. leia. The names of these and some other related beetles aim to grab your attention.
“We had wanted to do something fun like this for a while,” says Sarah Smith. She’s an insect biologist, or entomologist (En-toh-MOLL-uh-jist), at Michigan State University in East Lansing. The beetles’ features inspired their sci-fi names. C. leia, for example, is brown like Princess Leia’s buns. The pointy body shape of C. katniss reminded Smith of Katniss Everdeen and her bow and arrow.

Some names suit the beetles’ behavior too, says Anthony Cognato, another entomologist at Michigan State. He partnered with Smith on a new beetle study. Female ambrosia beetles venture out from nests to start families at new sites. This behavior may explain how these beetles spread throughout the tropics over the past 20 million years.
Their eye-catching names also draw attention to taxonomy. That’s the science of naming organisms. Taxonomy is a bedrock of biology, Cognato explains. A unique name lets scientists know which individuals fit within a species. Only then can they go on to discuss and study what makes this species different from others.
“This system of names provides the keys to unlock information,” adds Gustavo Hormiga. He’s at George Washington University in Washington, D.C. He studies spiders and other arachnids.
Imagine if you got a tick bite, Hormiga says. Some types of ticks carry Lyme disease, which causes rashes, fever and headaches. Other ticks don’t. A doctor deciding on treatment might ask if the bite was from a deer tick. This Ixodes scapularis can be a carrier of Lyme disease.
The way modern scientists classify and name organisms was first developed more than 250 years ago. But that doesn’t mean the process hasn’t changed. Since then, scientists have adapted the naming system to reflect how species evolve.
Some scientists, however, would like to start afresh with a new naming method. Many others are not ready for this. For most biologists, a name is not just a name. It’s also a way not only to distinguish what separates species, but also to understand species.
Linnaeus and his legacy
Modern taxonomy traces back to Carl Linnaeus (Leh-NAY-us). This Swiss naturalist wrote the book on it back in 1753. His Species Plantarum, as it was called, listed every plant species known at that time. It also marked the first time a binomial (two-part) nomenclature (naming system) came into common use for plants. Five years later, Linnaeus’s Systema Naturae used the same system to name animals. That book contained some now-familiar names, such as Homo sapiens and Boa constrictor.

Before binomial nomenclature, animal names involved long descriptions. Honeybees, for instance, were Apis pubescens, thorace subgriseo, abdomine fusco, pedibus posticus glabis, untrinque margine ciliates. This mouthful of Latin words describes the insects’ appearance. Soft short hairs, gray chest, brown belly and smooth back legs bordered with hairs on both sides. Linnaeus’s name for the species is much shorter: Apis mellifera, or “honey-bearing bee.”
The ordering of those two Latin words has a special meaning, too. The first describes the genus (GEE-nus). It’s the name for a related group. The second name specifies the particular species. The genus Canis, for instance, includes several species. They include the domestic dog (Canis familiaris) and the gray wolf (Canis lupus).
Modern scientists still use this binomial system, as when they named those sci-fi-inspired ambrosia beetles. The names also categorize organisms into groups, known as taxa. A group, or taxon (hence the field’s name: taxonomy), includes similar organisms. For example, the genus Canis makes up a taxon. The genus Vulpes, which includes foxes, makes up another taxon. Taxa (the plural form of taxon) also can describe larger categories, like the group of all mammals.
Linnaeus described five tiered rankings for animals. At the top were kingdoms. At the bottom were species. Over time, scientists added a few more ranks. The system now spans from kingdom down through phylum (FY-lum), class, order, family, genus and species. One way to keep track of these labels is to remember “King Phillip Came Over For Great Spaghetti.” The first letter of each word matches the first letter of a rank. Taxa can be defined at different ranks. For example, Mammalia (the group of all mammals) is a taxon at the rank of class.
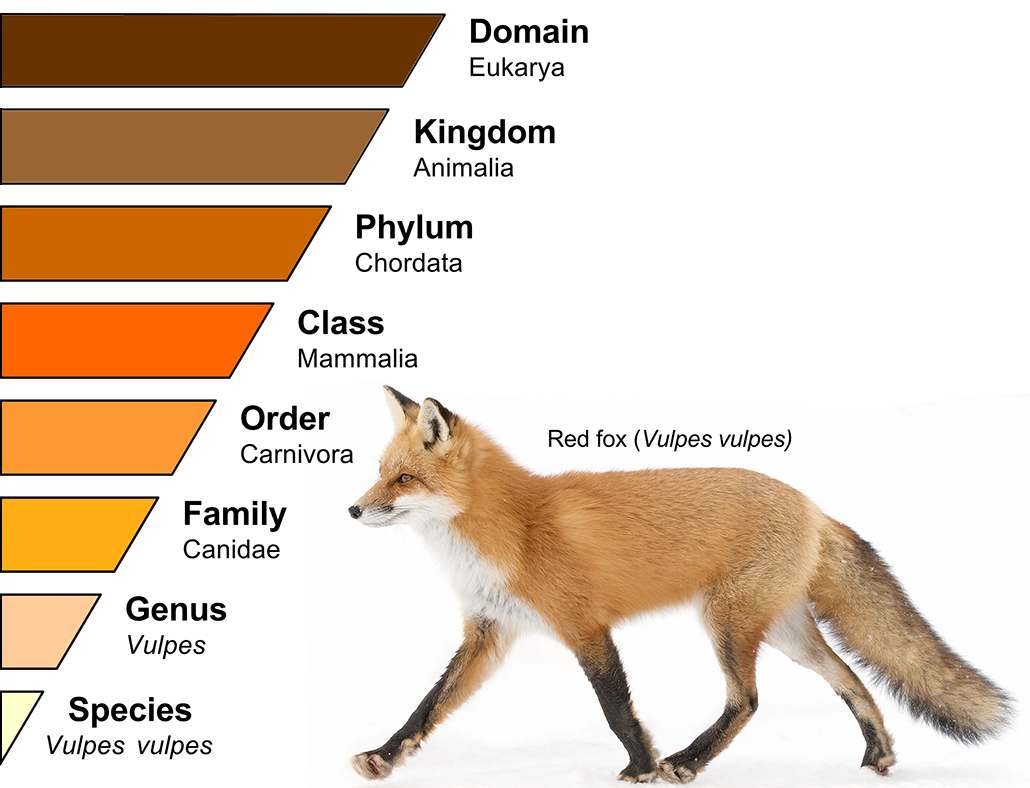
Remember how a genus can include multiple species? In the same way, an order can contain multiple families. And a class can contain multiple orders. The nested nature of these taxa provides scientists with helpful information.
“If I told you [two species] were in the same family, you would know right away that they were also in the same class and in the same phylum — and in the same kingdom,” explains Diana Lipscomb. She’s a retired evolutionary biologist at George Washington University.
However, there aren’t rules for deciding what rank a group of species — or taxon — falls into. One scientist might not agree with another about what defines a family, for example. In fact, a third scientist might even think the taxon should belong to a different rank, such as an order.
The International Code of Zoological Nomenclature, the rules for animal naming, spells out official guidelines. But this code doesn’t enforce rules for defining a taxon’s rank. That includes determining families versus orders. Because of this, Hormiga notes, all ranks may not be really comparable.
Evolving with the times
In the 1970s, scientists fiercely argued about taxonomy. These were turbulent years, Hormiga says. In the end, the field shifted. Scientists began to categorize species based on their evolution rather than on just what they looked like. This evolutionary approach is known as phylogeny (Fy-LAH-juh-nee).
“Modern taxonomy must reflect phylogeny,” says Gonzalo Giribet. He’s a zoologist at Harvard University in Cambridge, Mass. Members inside a taxon should be more related to each other than to species outside the group. Scientists draw these relationships as branched trees. In fact, Charles Darwin described such a tree of life. His included both living and extinct organisms.
I might tell you, Hormiga says, “there is a family of spiders unique to some part of South America” and it has been classified on the basis of phylogenetic relationships. “Then what I’m saying is, this is a branch of the tree of life.”
Taxonomy and phylogeny are key parts of a scientific field known as systematics. You might notice the name is related to Linnaeus’s Systema Naturae. This field uses evolution to classify and name living things.
Systematists (SIS-teh-muh-tists) study a species’ evolution by looking at its genetic material: DNA and RNA. DNA is a molecule that stores genetic instructions in cells. In order to read those instructions, cells copy DNA into another molecule, RNA. Cells use that RNA to make proteins.
Ribosomes (RY-boh-soams) are the cellular machines that make those proteins. Each ribosome is made of proteins and RNA. Because ribosomes are so important to cells, changes in RNA sequences could prove a disaster. So mutations — copying changes — rarely happen in ribosomal RNA. Over millions and billions of years, however, a few changes can slowly accumulate as species evolve. The more different the RNA in two species is, the less related those species will be.
Scientists have used such analyses to divide life on Earth into three domains. Two describe much of all one-celled life: bacteria and archaea (Ar-KEE-ah). The third — eukaryotes (Yu-KAIR-ee-oats) — includes everything else. Eukaryotes can be one or more cells. All have cellular parts with distinct roles, such as DNA-containing nuclei. Plants fit in this group. So do all animals. So, too, do one-celled amoebas.
You can forgive Linnaeus for not coming up with this system. He lived long before scientists grasped the idea of evolution, much less genetics. As a result, new findings can force systematists to re-classify taxa. For example, Smith and Cognato moved some ambrosia beetle species into the genus Coptoborus. This was after they discovered that Coptoborous and the genus Theoborus were actually one and the same.
To many systematists, these changes are okay. “You’re making a hypothesis about what this organism is, who it’s related to and where it fits in on that tree of life,” says Lipscomb at George Washington. A big part of science is interpreting new data to improve our understanding of our world (and cosmos).
Bigger than kingdoms, perhaps?
Domains are just one update in taxonomy’s highest ranks. In Systema Naturae, Linnaeus described three living kingdoms: animals, plants and minerals. It’s no surprise that the mineral kingdom was jettisoned. Since then, scientists have defined more kingdoms. One popular model includes five: Animalia, Plantae, Fungi, Protista and Monera.
Kingdom Monera includes prokaryotes (Pro-KAIR-ee-oats). These are single-celled organisms without a nucleus. So it includes bacteria and archaea. The other four kingdoms cover eukaryotes. So you can likely guess what Animalia, Plantae and Fungi include. Kingdom Protista becomes a grab-bag for anything that’s not an animal, plant or fungus but still has cells with nuclei.
In terms of evolution, that last group has a problem. Some of its members are more related to animals or plants than to other protists.
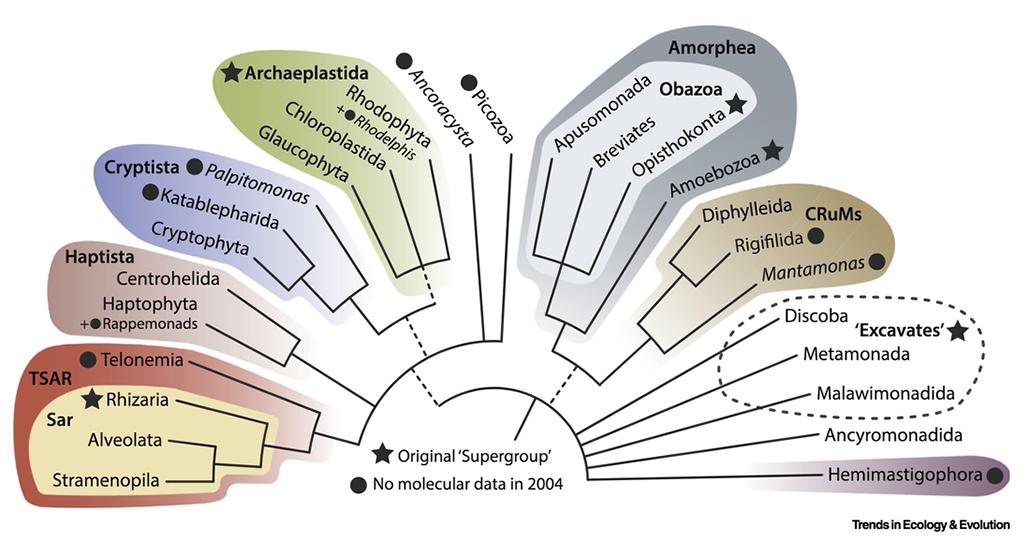
Researchers studying single-celled organisms don’t consider Protista a true taxon, says Alastair Simpson. He’s a biologist at Dalhousie University in Halifax, Canada. Instead, he says, it’s more helpful to talk about supergroups. These divide eukaryotes into about a dozen groups based on how they evolved.
Simpson was a co-creator of the supergroup concept in the early 2000s. The choice of the name “supergroup” was intentional. “We deliberately didn’t want something that sounded like a rank,” Simpson says. “We thought supergroups would be silly enough.”
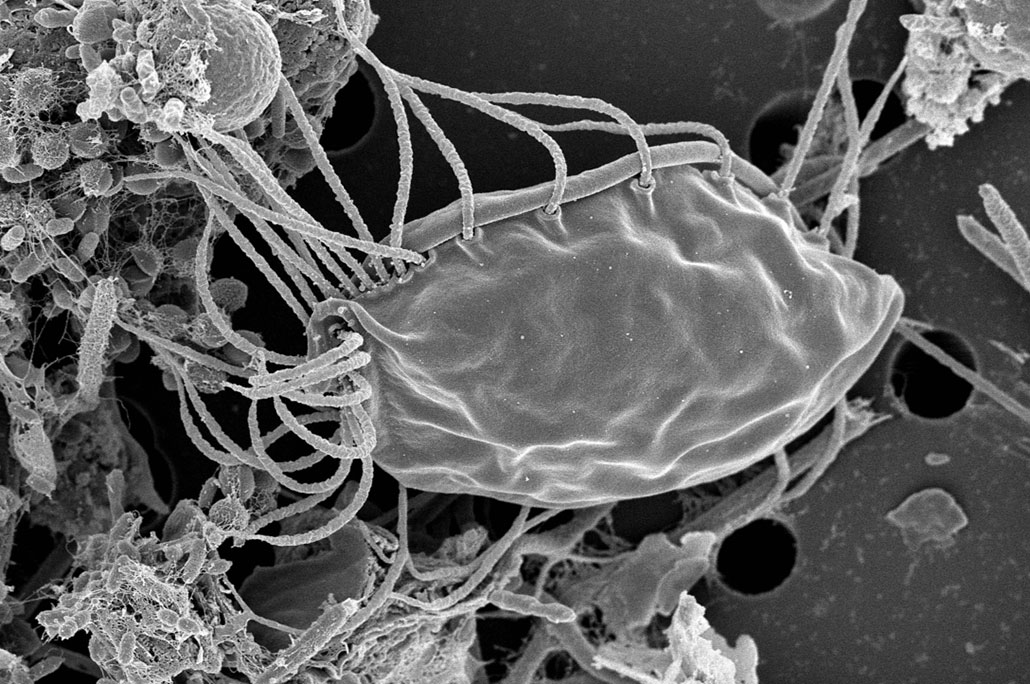
Animals and fungi fall under one supergroup: It’s called Obazoa. Other supergroups highlight the diversity of single-celled eukaryotes once lumped together. Simpson coauthored a recent paper describing a new single-celled species, Hemimastix kukwesjijk. Its name comes from the traditions of the Mi’kmaq First Nation of Nova Scotia, Canada. In their Indigenous language, the name translates as “little ogre.”
These microbes “are evolutionarily distant from everything else that we know about,” Simpson says. As a result, he and his colleagues believe that the phylum containing the new species actually makes up a supergroup — a category bigger than a kingdom.
Should biologists start over from scratch?
Some scientists have been developing a new system for naming species. “We’ve developed this alternative approach” so names wouldn’t change as much as in a rank-based system, explains Kevin de Queiroz. A zoologist, he is the curator of reptiles and amphibians at the National Museum of Natural History. It’s part of the Smithsonian Institution in Washington, D.C.
The new naming system is called PhyloCode. That’s short for International Code of Phylogenetic Nomenclature. As its name implies, species names are tied to how strongly they are linked through evolution, says de Queiroz. He’s a PhyloCode developer. And although many people may not have heard of the PhyloCode, de Queiroz is part of a team that has spent the last 20 years getting it ready for prime time. PhyloCode’s sixth edition was published in 2020. It was the first to be published as a printed book. A companion volume, Phylonyms, provides details about how to define groups.
How does it work? With the current naming system, the name “Iguanidae” is the family (based on the -idae ending) containing the genus Iguana. But remember, scientists decide ranks for taxa. If a scientist decides “Iguana” should apply to a bigger or smaller group, the update could cause names to shift for other taxa and species.
PhyloCode instead defines names based on how species are linked by evolution. A new version of Iguanidae is defined as a specific ancestor of the green iguana (Iguana iguana) and all its descendants. You can think of this as a branch on the tree of life. This branch that defines Iguanidae excludes the common agama and chameleon.
These new types of definitions no longer rely on ranks, although they don’t necessarily throw them out. Ranks can still provide information about how groups relate to each other on the tree of life. There’s no problem with using ranks to show how mammals (a class) are part of a larger branch of animals (a kingdom), for example. “But you shouldn’t be using [ranks] to determine how the names are applied,” de Queiroz argues.
This proposal to change the naming system for all living things has produced a range of responses. There are boosters like de Queiroz. Many outright oppose it. Others fall somewhere in between.
“I agree with [de Queiroz] in principle,” says herpetologist Darryl Frost. Yet Frost is not completely on board. “Do I have any issues with them in terms of practicality,” he asks? “Yeah, I do.” Frost studies reptiles and amphibians at the American Museum of Natural History in New York City.
And he isn’t alone in having concerns. “PhyloCode is not a very widely accepted concept,” Simpson says. He’s a contributor to Phylonyms and has written chapters about protists. One problem with the new system, he says, is that a name can stay the same while the ideas behind it changes. Time will tell whether biologists come to fully embrace PhyloCode — and how it integrates existing names or naming systems.
Taxonomy and biodiversity
New developments in taxonomy are more important than ever. Humans have made a big impact on the Earth’s multitude of species — and generally not in a good way.
“Species are going extinct at a tremendous rate,” Hormiga says. “Once they’re extinct, they’re gone forever.” Unfortunately, there’s a shortage of trained taxonomists to describe every living thing on the planet that has yet to be named.
This presents a huge challenge, as taxonomy is crucial to slowing the biodiversity crisis. Why? Explains Frost, “You have to name it if you’re going to conserve it.”
https://www.sciencenewsforstudents.org/article/biology-species-name-linnaeus-taxonomy